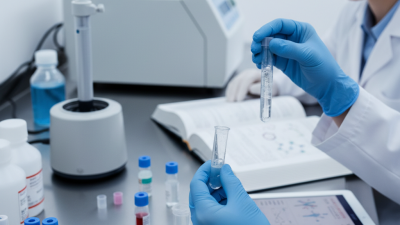
Genomic DNA Cleanup is an essential step in molecular biology. It ensures the purity and integrity of DNA for various applications. Researchers need effective methods to refine their genomic samples.
There are numerous techniques available for Genomic DNA Cleanup. Some methods may be more suitable for certain types of samples. Others may come with specific limitations that researchers must consider. For example, purification kits are convenient, but they can be costly. On the other hand, traditional methods, while economical, require careful execution to avoid contamination.
As we explore the top ten methods for Genomic DNA Cleanup, it’s crucial to understand their pros and cons. Each method has its unique challenges and benefits. Selecting the right approach may involve trial and error. However, informed choices lead to successful results. Understanding the nuances of different techniques will enhance your research outcomes. Ultimately, the goal is to achieve high-quality genomic DNA for reliable results.

Genomic DNA cleanup is crucial for accurate results in molecular biology. Several techniques exist for this process. However, understanding the essential principles can greatly improve your outcomes. One key principle is the minimization of contamination. Always use clean tools and reagents to avoid introducing foreign DNA. This simple practice can save time and resources.
Another important aspect is choosing the right method based on your sample type. Different techniques, such as silica membrane-based purification, target specific contaminants effectively. However, they may also have limitations. Sometimes, they don't perform well with certain samples or conditions. Acknowledging these limitations helps refine your approach.
The final principle is the importance of thoroughness. It's easy to overlook steps, but this can compromise the DNA quality. Each step in the cleanup process, from binding to elution, matters. Missing even a small detail can lead to unwanted results. Take the time to review your process and reflect on areas for improvement. This reflection fosters growth and enhances your skills in genomic DNA cleanup.
Genomic DNA extraction often encounters several common contaminants. These include proteins, phenols, and salts. Removal of these impurities is crucial for successful downstream applications. Protein contamination can interfere with enzymatic reactions. A study by the National Institutes of Health emphasizes that nearly 30% of samples may retain significant protein contaminants after standard extraction procedures.
Phenolic compounds, often from plant materials, pose another challenge. They can inhibit PCR reactions and affect sequencing quality. Data from the Journal of Molecular Biology suggests that around 40% of extractions from complex tissues may have phenolic contamination. Salt impurities can also affect the quality of genomic DNA. High salt concentrations may cause precipitation issues during further analyses. It's not uncommon for researchers to overlook these contaminants.
Despite all precautions, some inherent issues remain. For instance, genomic DNA extraction efficiency varies considerably across tissue types. Sometimes, contaminants slip through meticulous cleanup processes. Continued improvements are needed. Recognizing specific contaminants is crucial to enhance extraction protocols. Addressing these issues can lead to more reliable genomic studies.
Effective genomic DNA cleanup is crucial in molecular biology. It ensures the purity and integrity of DNA samples for accurate results. Several methods are available, each with its own advantages. Common techniques include phenol-chloroform extraction and silica-based methods. These techniques are widely used due to their effectiveness and reliability.
One method worth considering is enzymatic cleanup. It employs specific enzymes to digest unwanted materials. This approach can be time-consuming but often yields high-quality DNA. Another approach is magnetic bead-based cleanup. This method uses magnetic particles to isolate DNA easily. It is efficient and reduces the risk of contamination.
However, it’s essential to evaluate each method's protocols carefully. Some techniques may be more suited to specific applications or sample types. It’s also important to consider the trade-offs, such as time versus yield. Not every method will work flawlessly every time. Users should be prepared to troubleshoot and adapt their methods. Effective cleanup can lead to reproducible experimental outcomes, but imperfection is always possible.
When conducting genomic DNA cleanup, it's vital to evaluate the efficiency and yield of various techniques. Different methods can significantly impact the purity of DNA samples. Some techniques are more favorable for certain applications due to their unique properties. For instance, spin-column methods often yield high purity but may have lower overall recovery rates.
In contrast, magnetic bead-based cleanup offers a versatile approach. This method can efficiently capture DNA in varying lengths. However, researchers sometimes find it challenging to optimize the binding and washing conditions. Such inconsistencies can lead to substantial variations in yield. Additionally, precipitation methods can be effective but require careful handling. Even slight deviations in temperature or time can affect results.
Understanding these nuances is essential. The goal is to find a balance between efficiency and yield. Experimenting with several techniques can reveal their strengths and weaknesses. Each method has its conditions and limitations. Awareness of these factors allows for better decision-making in genomic cleanup processes.

Assessing DNA quality after cleanup is vital for accurate results in genomics. High-quality genomic DNA is essential for downstream applications, including sequencing and genotyping. Key indicators of DNA integrity include the A260/A280 ratio and gel electrophoresis results. A common benchmark is achieving a ratio between 1.8 and 2.0 for pure DNA. Deviations from this range often indicate contamination or degradation.
Recent studies highlight that over 30% of genomic samples can yield suboptimal quality post-cleanup. This rate is concerning for researchers who rely on precise outcomes. Regardless of the cleanup method used, a thorough quality assessment must follow. Techniques like Nanodrop and Qubit offer quick quantitative measures. Additionally, assessing fragment size distribution using a Bioanalyzer can provide insights into DNA fragmentation.
Some researchers may overlook RNA contamination, which can lead to erroneous interpretations. Ensuring that the DNA sample is free from impurities is key. Reviewing batch effects is essential for reproducibility and reliability. Awareness of potential pitfalls in the cleanup process will improve overall data accuracy and integrity.