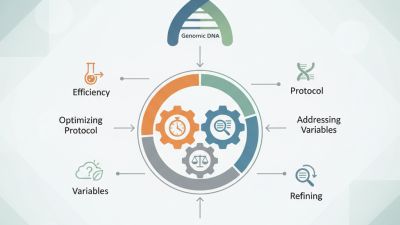
In preparation for the upcoming 139th Canton Fair, where advanced technologies and manufacturing solutions will be showcased, the importance of effective DNA Library Preparation for NGS cannot be overstated. According to a recent industry report by Genomic Innovations, the demand for next-generation sequencing methods has grown exponentially, with a projected increase of 25% annually. This highlights a significant opportunity for industries centering on genomic research and applications.
Dr. Emily Carter, a recognized expert in genomic sequencing, states, "The efficiency of DNA Library Preparation for NGS can directly impact research outcomes." This underscores the need for high-quality preparation techniques, as improper handling leads to data loss. Furthermore, advancements such as AI in supplier search at the Canton Fair may aid in identifying leading providers of genomic solutions.
Yet, challenges remain within the field. Many labs struggle with standardizing protocols, which can yield inconsistent results. These discrepancies often stem from inadequate training and resource allocation. Therefore, it's crucial to focus on quality methodologies for DNA Library Preparation for NGS to ensure robust and reproducible results in genetic research.

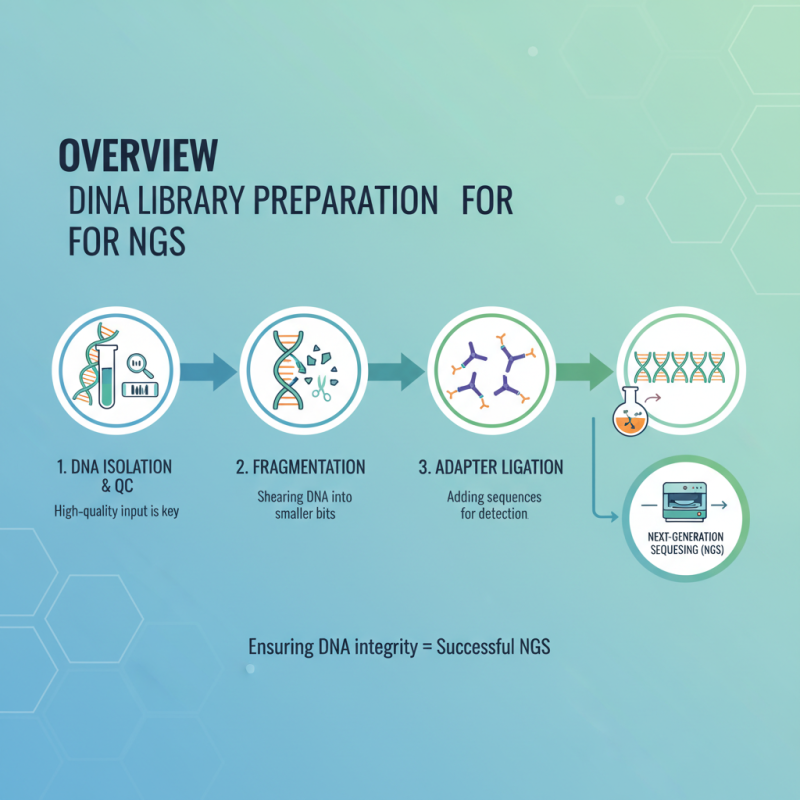
DNA library preparation is a crucial step for next-generation sequencing (NGS). This process transforms fragmented DNA into a form suitable for sequencing machines. It begins with the isolation of high-quality DNA. The quality of the input material significantly affects the outcome. Researchers often face challenges in maintaining the integrity of DNA samples.
Once DNA is isolated, it undergoes fragmentation. This can be achieved through enzymatic digestion or mechanical shearing. Fragment sizes need to be carefully controlled. If fragments are too long or too short, the final library may not perform well. Aiming for optimal size is vital. After fragmentation, adapters are added to the DNA fragments. These adapters help the sequencer bind and process the DNA. However, sometimes, the efficiency of adapter ligation can vary. Troubleshooting this step is often necessary.
The final stage involves amplifying the library. This is done using polymerase chain reaction (PCR). While PCR can enhance yield, it may introduce biases. Over-amplification can distort the representation of certain sequences. It is important to assess the library quality before sequencing. Analyzing size distribution and concentration ensures that the library is ready for NGS. These steps may seem straightforward, but achieving consistent results requires practice and attention to detail.
Preparing a DNA library for NGS requires specific components and materials. Start with high-quality genomic DNA. Ensure it is well purified and free from contaminants. Fragmentation of the DNA is crucial. Use enzymatic or mechanical methods to achieve the desired size. A small fragment size increases efficiency in sequencing.
Adapters are essential in this process. They link to the DNA fragments, providing binding sites for sequencing. Choose adapters that suit your sequencing technology. Another important element is the DNA polymerase. This enzyme amplifies the DNA library. A good quality polymerase ensures accurate and high yields.
Tips: Always verify your DNA concentration. Use a fluorometer for precise measurements. This avoids issues during library construction. Make sure to run a quality control step after each major phase. This helps identify potential problems early on. Take notes on your workflow. It may highlight moments that need improvement for future runs.

Preparing a DNA library for next-generation sequencing (NGS) involves several essential steps. Initially, you'll start with the extraction of high-quality DNA. Fragment the DNA into smaller pieces. This process can be imprecise; ensuring uniform sizes is often a challenge. An optimal size range is crucial for efficient sequencing.
Next, adaptors must be ligated to the DNA fragments. The choice of adaptors can significantly influence the results. It’s important to test different adaptor types. Some might not bind well, leading to wasted samples. Following this, amplify the library using PCR. This step is vital but can also introduce bias. Monitoring the PCR cycles is necessary to avoid over-amplification.
Lastly, purify the library to eliminate excess reagents. This step can be tricky, as residual components may interfere with sequencing. Quality control is essential here; running a gel electrophoresis can help visualize the results. Mistakes in this phase can affect the entire project. Each step requires careful attention, as small errors can lead to significant setbacks down the line.
Quality control is crucial in DNA library preparation, especially for next-generation sequencing (NGS). A recent report published by the National Center for Biotechnology Information states that approximately 20% of NGS libraries fail standard quality assessments. This underscores the importance of rigorous QC measures. Inspecting the size distribution of DNA fragments is one way to ensure the integrity and usability of your library. Using bioanalyzers or gel electrophoresis can help identify any size discrepancies.
Another major aspect involves quantifying DNA concentration accurately. It’s essential to obtain a precise measurement. The inaccuracies can lead to either oversaturation or low yield during sequencing. The ideal concentration should typically range from 2 to 4 nM for most sequencing platforms. However, a significant number of users report struggles with achieving this concentration, pointing to potential procedural flaws. These discrepancies often stem from inefficient purification methods or even the use of degraded reagents.
Amplicon libraries require additional attention. Despite their popularity, they can often show unexpected variations in performance. This inconsistency can result from various factors, including PCR biases. Many scientists overlook the need for multiple rounds of validation to detect these biases early. It’s essential to rethink standard practices, emphasizing comprehensive QC rather than relying solely on initial metrics. Quality control measures deserve focused improvement to enhance library preparation for NGS.
| Step | Description | Quality Control Measure | Expected Outcome |
|---|---|---|---|
| 1 | DNA Extraction | A260/A280 Ratio | 1.8 - 2.0 |
| 2 | Fragmentation | Average Fragment Size | 200 - 800 bp |
| 3 | End Repair | Quality Check on Bioanalyzer | High Yield |
| 4 | Adapter Ligation | Quantification via qPCR | Optimal Library Concentration |
| 5 | Cleanup | Nanodrop Quantification | Concentration within standards |
Preparing a DNA library for next-generation sequencing (NGS) requires careful attention to detail. Start with high-quality DNA samples. Check the concentration and purity thoroughly. Use a spectrophotometer or fluorometer for accurate measurements. Avoid contamination to ensure valid results. It’s wise to run a gel electrophoresis to check fragment size. This step can often reveal inconsistencies in sample quality.
Consider choosing the right library preparation kit. Different projects may need different methods. Follow the manufacturer's protocol closely but be open to adjustments if necessary. It’s common to encounter issues like unexpected band sizes on the gel. Reflect on those moments and tweak your approach. One small error in the pipetting process can lead to major discrepancies.
During the preparation, document every step meticulously. Keeping track of your process aids in troubleshooting later. Mistakes happen; it's part of science. Even with the best preparation, results can still be unpredictable. Embrace these challenges as learning opportunities. Each session at the lab can improve your skills. The journey of mastering NGS library preparation is ongoing.