Genomic DNA cleanup is a crucial step in molecular biology that ensures the purity and integrity of DNA samples. According to a report by the National Center for Biotechnology Information, over 70% of researchers encounter challenges in obtaining high-quality genomic DNA. These challenges can lead to inconsistent results in downstream applications, emphasizing the need for effective cleanup techniques. Dr. Emily Carter, a leading expert in the field, once stated, “The quality of DNA cleanup directly impacts the success of your experiments.”
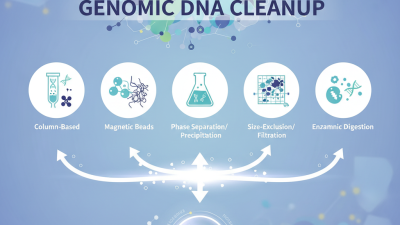
When performing Genomic DNA cleanup, various methods exist, including column-based purification and magnetic bead-based approaches. Each method has its pros and cons, which can complicate the decision-making process for researchers. Recognizing the specific needs and constraints of your experiment is essential. Some purification methods may be quick but can leave impurities that influence the experimental outcomes. Effective cleanup requires not just the right technique but also an understanding of the nuances involved in sample handling.
Moving forward, researchers must reflect on their methods and adapt them to their ever-evolving needs. Commitment to high standards in Genomic DNA cleanup will ultimately enhance the reliability of research findings. It’s crucial to assess methods and results systematically, as even minor oversights can lead to significant consequences in the lab.

Genomic DNA cleanup is a critical step in molecular biology. It ensures that the DNA sample is free from contaminants such as proteins, solvents, and enzymes. These substances can interfere with downstream applications like PCR and sequencing. Purifying DNA leads to more reliable and reproducible results in scientific experiments. It is essential for obtaining high-quality data, especially in research settings.
Tips: Always use fresh reagents. Contaminated or expired chemicals can compromise your cleanup. Pay attention to handling practices, as cross-contamination can occur easily. Use designated pipettes and tips for genomic DNA to maintain sample integrity.
Cleaning up genomic DNA can be complex. Many researchers struggle with achieving complete purity and yield. It’s not unusual to observe variations in DNA concentration. Sometimes, it might take multiple cleanup attempts to get satisfactory results. Be prepared to analyze your methods critically and adjust where necessary.
Tips: Run a gel electrophoresis test after cleanup. This helps visualize DNA integrity. If results are not ideal, tweak your protocol. Evaluate the impact of each step on the final outcome, and be open to trial and error.
Genomic DNA cleanup is a crucial step in molecular biology. Properly cleaning up DNA ensures that it is free from contaminants and inhibitors. These impurities can affect downstream applications, like sequencing and cloning. Key steps in the genomic DNA cleanup process focus on purification and concentration.
The first step involves cell lysis. It breaks open cells to release DNA. This is often achieved using buffers that disrupt cell membranes. After lysis, proteins and other debris must be removed. This can be done through centrifugation or filtration. However, it's essential to assess whether complete removal has occurred. Undetected proteins can interfere with results later.
Another vital step is precipitation. Alcohol is typically used to precipitate DNA. It’s key to carefully control the temperature and time during this process. Temperature fluctuations can lead to inconsistent outcomes. Finally, washing the DNA pellet requires attention. Ethanol washes help remove residual salts. Again, this step can be overlooked. Incomplete washing can lead to downstream complications. Each step in the cleanup process demands precision. Any oversight could undermine the quality of the genomic DNA.
When performing genomic DNA cleanup, the choice of tools and reagents significantly impacts the efficiency of the process. Commonly used reagents include ethanol, sodium acetate, and various silica-based columns. According to a report by Nature, using a silica membrane can enhance DNA yield by up to 30% compared to traditional methods. This emphasizes the importance of selecting appropriate materials for optimal results.
Standardizing protocols is crucial to achieving consistent outcomes. Many labs report variability due to differences in reagent quality and handling. A study in the Journal of Molecular Biology found that inconsistencies led to a 15% decrease in DNA quantity. Adopting standardized practices ensures reliability and minimizes variability.
Additionally, the technique of binding and washing DNA needs careful attention. Excessive washing can lead to DNA loss. Researchers often overlook this, assuming more washing equates to purer DNA. In reality, over-washing may inadvertently remove smaller fragments. Balancing thoroughness with caution is essential in the cleanup process. Proper tool selection, along with a mindful approach to reagent usage, can significantly improve genomic DNA cleanup efforts.
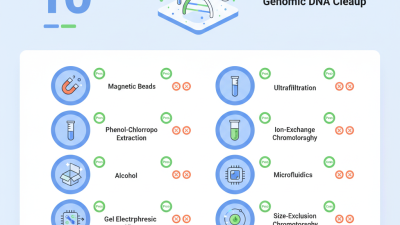
Genomic DNA cleanup is crucial for accurate downstream applications. However, it comes with several challenges. One frequent issue is the presence of contaminants. Particles like proteins, RNA, and salts may co-purify with DNA, leading to unreliable results. Studies indicate that 25% of low-yield DNA samples suffer from such contaminants. Effective methods to handle this include using variations of phenol-chloroform extraction. This method enhances purity but may introduce toxic substances, demanding careful handling.
Another challenge is the inefficiency of traditional cleanup protocols. They often require multiple steps that increase the risk of error. Around 40% of researchers report inconsistencies in their cleaning processes. To mitigate this, incorporating magnetic bead-based techniques has shown promise. Magnetic beads provide a more uniform purification process. They allow for easy separation and reduce the risk of DNA loss. This method can boost yield consistency, yet researchers must remain vigilant about bead-to-sample ratios.
Temperature and time also play pivotal roles in DNA cleanup. Incubation at incorrect temperatures may degrade DNA quality. Data suggests that 30% of samples could be compromised due to improper handling. When using enzymatic cleanup methods, precise timing is vital. Experimenting with variables can lead to improvements, but it requires a willingness to embrace trial and error.
| Challenge | Description | Solution | Best Practices |
|---|---|---|---|
| Low DNA Yield | Insufficient recovery of DNA during purification. | Optimize lysis buffer and increase incubation time. | Ensure proper sample mixing and avoid excessive centrifugation. |
| Contamination | Presence of RNA or proteins in the purified DNA. | Use DNase and protease during cleanup. | Use filter tips and work in a clean area to avoid cross-contamination. |
| Fragmentation | DNA samples become fragmented during handling. | Implement gentle handling and avoid freeze-thaw cycles. | Use low-speed centrifugation and keep samples cold during processing. |
| Inefficient Cleanup | Inadequate removal of impurities. | Use a combination of silica columns and magnetic beads. | Follow the protocol precisely and add appropriate wash steps. |
| Storage Issues | Degradation of DNA during storage. | Store at -20°C or -80°C in appropriate buffers. | Use aliquots to prevent repeated freeze-thaw cycles. |
Quality control after genomic DNA cleanup is crucial for reliable results. Contamination or degradation can lead to inaccurate conclusions. Studies show that about 30% of DNA samples may become compromised during cleanup processes. Therefore, it is essential to implement effective quality control measures.
One effective practice involves quantifying DNA using fluorometric methods. These methods can detect low concentrations accurately, which is vital when working with minimal samples. A study published in *Nature* revealed that fluorometry can double the accuracy of DNA quantification compared to spectrophotometric methods. Additionally, checking DNA purity with the A260/A280 ratios can highlight potential contaminants, but it may not fully detect all impurities.
Another important aspect is integrity assessment. Running DNA samples on an agarose gel helps visualize fragmentation. This step ensures that the DNA has maintained its structural integrity. Some researchers overlook this step, mistakenly believing that cleanup methods alone guarantee quality. Committing to these quality control practices can significantly enhance the reliability of genomic analyses and the overall validity of research findings.